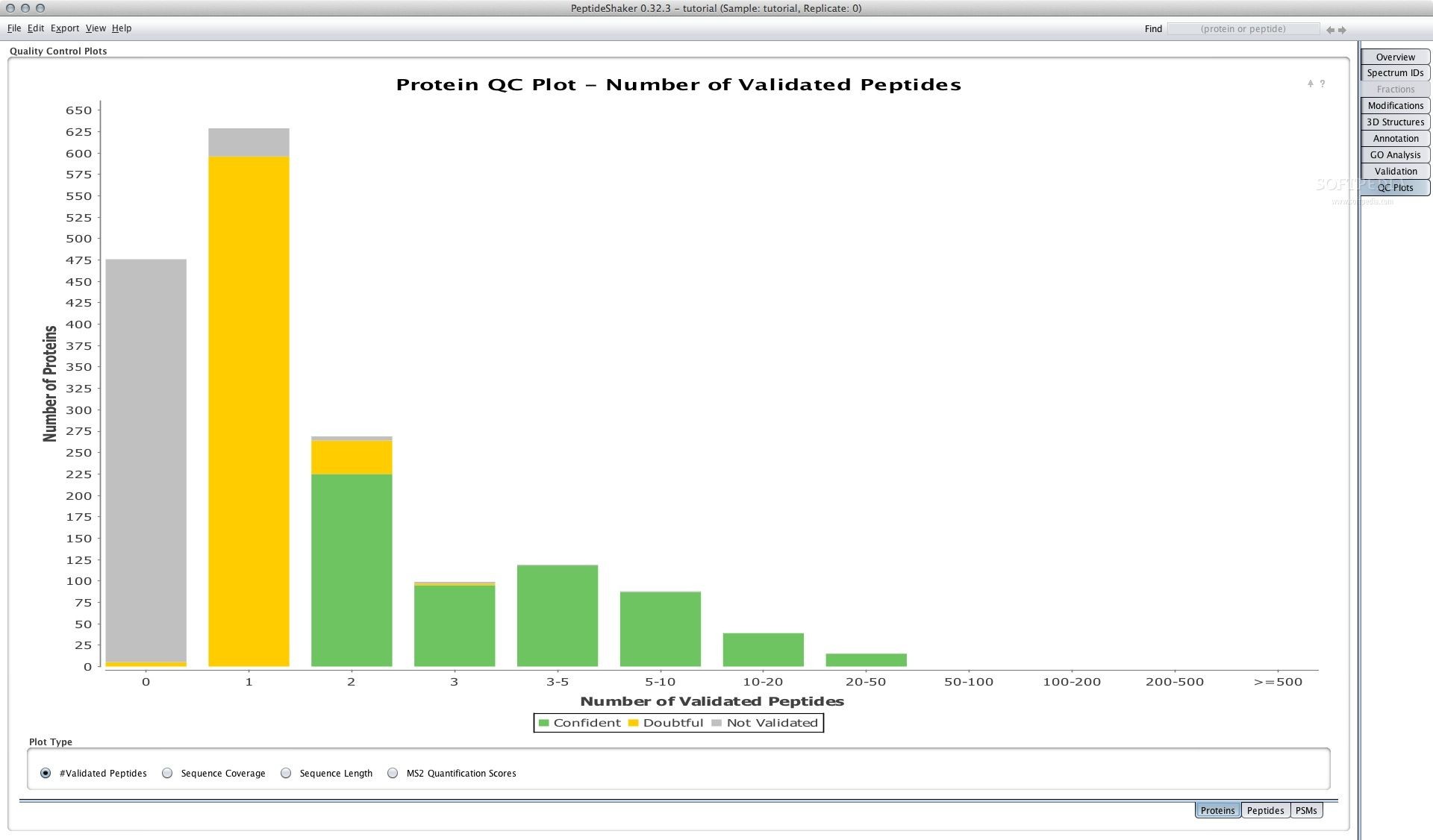
The program queries multiple online engines for complete protein and peptide analyses and statisticsĪll in all, PeptideShaker is a great tool for scientists and other professionals involved in studying proteins and peptides. When cases allow it, individual strings are highlighted in different colors, allowing for quick comparisons between data sets. Where applicable, the application generates data-rich graphics in multiple forms, including bar charts and bubble plots. PeptideShaker provides users with a wealth of statistical and descriptive data, including the number of peptides, spectrum matches sequences and gene ontology enrichment analyzes. 
By combining the results from multiple search engines, while re-calculating PTM localization scores and redoing the protein inference. These can be accessed from the right-side panel. PeptideShaker is a search engine independent platform for interpretation of proteomics identification results from multiple search engines, currently supporting XTandem, MS-GF+, MS Amanda, OMSSA, MyriMatch, Comet, Tide, Mascot, Andromeda and mzIdentML. There are nine analysis tasks supported, ranging from overviews to 3D structures and QC plots. These technical specs left to the side, the application provides users with a comprehensive protein analysis environment. Generate informative and detailed diagrams Also, the latest version of Java should be installed (if one cannot launch the program, users should try the 圆4 bit version). interpretation of proteomics identification results. However, before any proteins are processed, users should know that the application is highly resource-intensive and requires at least 4 GB of RAM. This diminishes the risk of errors and ensures data is properly analyzed. With it, users can query nine platforms for the interpretation of Proteomics results: X!Tandem, MS-GF+, MS Amanda, OMSSA, MyriMatch, Comet, Tide, Mascot, and mzIdentML. PeptideShaker is one such tool that goes about the problem in a different method. 
PeptideShaker is a search engine independent platform for the interpretation of proteomics identification results from multiple search engines. Fortunately, the exponential increase in digital tools has also meant that biologists, chemists, physicists and all other professionals interested in the field of Proteomics now have powerful analysis tools at their disposal. Interpretation of proteomics identifications from multiple search engines.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
